Pattern simulation
Functions used to simulate localization patterns.
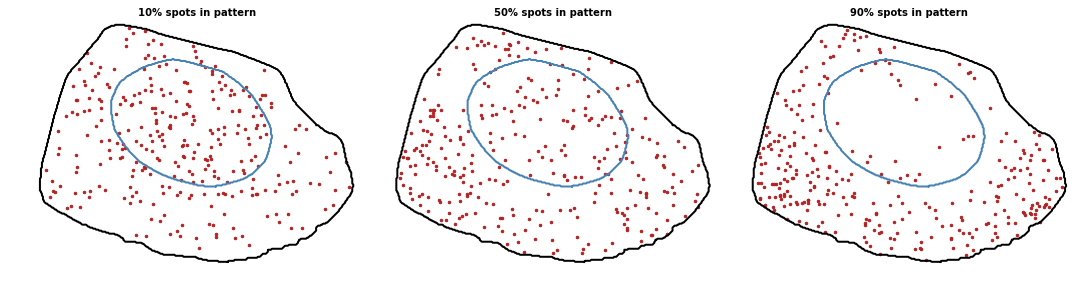
We implemented 9 patterns (8 localized patterns + 1 default random pattern) in 3D. The percentage of localized spots defines the pattern strength:
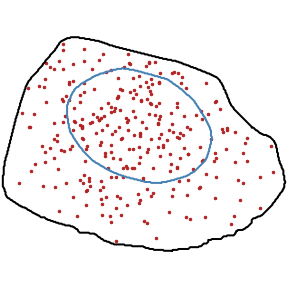
Random
Foci
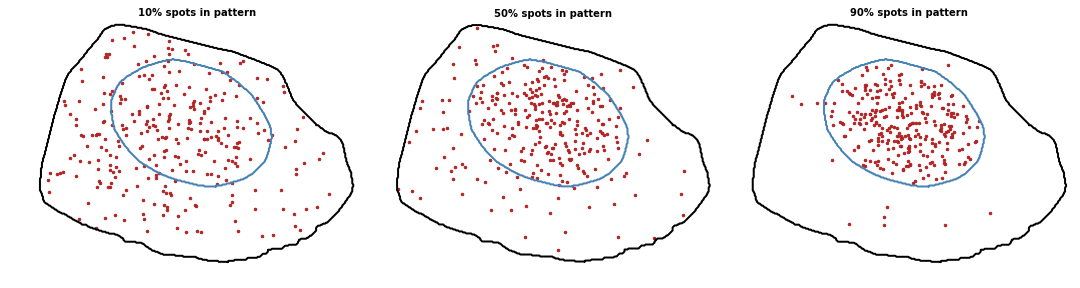
Intranuclear
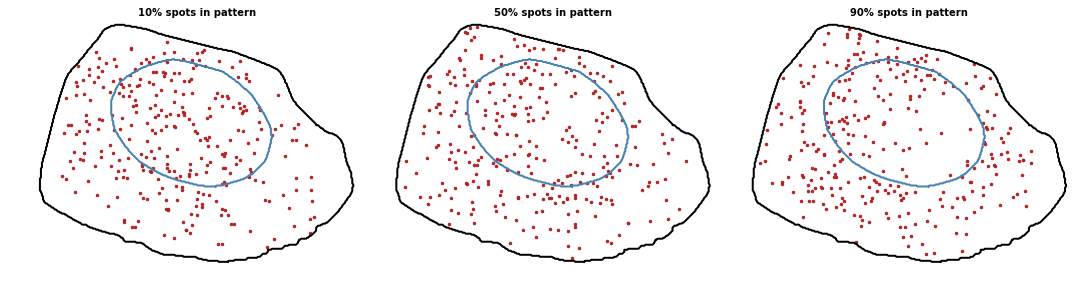
Extranuclear
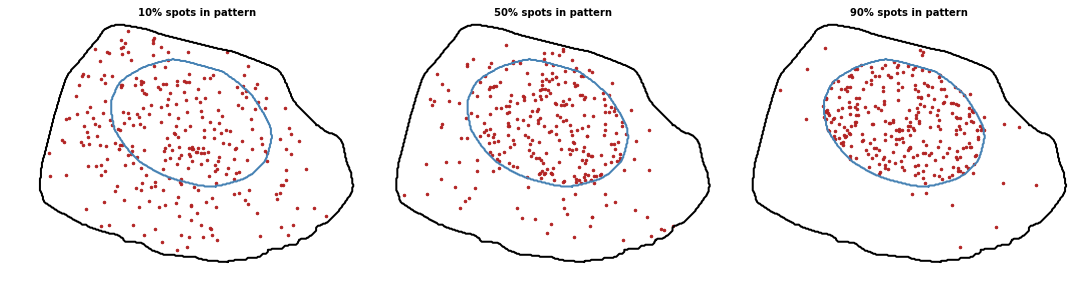
Nuclear edge
Perinuclear
Cell edge
Pericellular
Protrusion
We build a map of probability distribution to bias the localization of generated spots. Maps are built from specific cell templates:
We can simulate ground truth coordinates based on these probability maps:
- simfish.build_probability_map(path_template_directory, i_cell=None, index_template=None, map_distribution='random', return_masks=False)
Build a template and its probability map to sample spot coordinates.
- Parameters
- path_template_directorystr
Path of the templates directory.
- i_cellint, optional
Template id to build (between 0 and 317). If None, a random template is built.
- index_templatepd.DataFrame, optional
Dataframe with the templates metadata. If None, dataframe is load from ‘path_template_directory’. Columns are:
‘id’ instance id.
‘shape’ shape of the cell image (with the format ‘{z}_{y}_{x}’).
‘protrusion_flag’ presence or not of protrusion in the instance.
- map_distributionstr, default=’random’
Probability distribution map to generate among ‘random’, ‘random_out’, ‘random_in’, ‘nuclear_edge’, ‘perinuclear’, ‘cell_edge’, ‘pericellular’ and ‘protrusion’.
- return_masksbool, default=False
Return cell and nucleus binary masks.
- Returns
- probability_mapnp.ndarray, np.float32
Probability map to sample spots coordinates with shape (z, y, x).
- cell_masknp.ndarray, bool, optional
Binary mask of the cell surface with shape (z, y, x).
- nuc_masknp.ndarray, bool, optional
Binary mask of the nucleus surface with shape (z, y, x).
- simfish.simulate_localization_pattern(path_template_directory, n_spots, i_cell=None, index_template=None, pattern='random', proportion_pattern=0.5)
Simulate spot coordinates with a specific localization pattern from a cell template.
- Parameters
- path_template_directorystr
Path of the templates directory.
- n_spotsint
Number of spots to simulate.
- i_cellint, optional
Template id to build (between 0 and 317). If None, a random template is built.
- index_templatepd.DataFrame, optional
Dataframe with the templates metadata. If None, dataframe is load from ‘path_template_directory’. Columns are:
‘id’ instance id.
‘shape’ shape of the cell image (with the format ‘{z}_{y}_{x}’).
‘protrusion_flag’ presence or not of protrusion in the instance.
- patternstr, default=’random’
Spot localization pattern to simulate among ‘random’, ‘foci’, ‘intranuclear’, ‘extranuclear’, ‘nuclear_edge’, ‘perinuclear’, ‘cell_edge’, ‘pericellular’ and ‘protrusion’.
- proportion_patternint or float, default=0.5
Proportion of spots to localize with the pattern. Value between 0 and 1 (0 means random pattern and 1 a perfect pattern).
- Returns
- instance_coorddict
Dictionary with information about the cell:
cell_id: Unique id of the cell.
bbox: bounding box coordinates with the order (min_y, min_x, max_y, max_x).
cell_coord: boundary coordinates of the cell.
cell_mask: mask of the cell.
nuc_coord: boundary coordinates of the nucleus.
nuc_mask: mask of the nucleus.
rna_coord: rna spot coordinates.